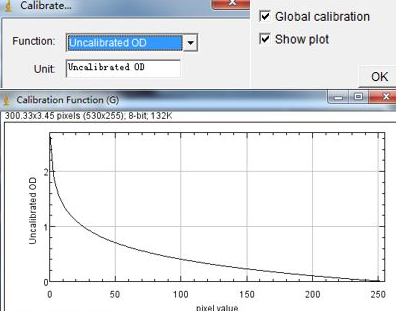
This is accomplished through the Skeletonize (2D/3D) plugin's methods and has the added benefit of operating on 3D datasets as well as 2D. The first, iterative thinning, is the method used in the original macros and produces a skeleton by iteratively removing outer pixels until a one pixel wide structure remains. The skeleton itself can be generated in two ways. The simplification is the generation of a morphological skeleton from which polylines can be extracted and analyzed (as is accomplished by the Analyze Skeleton plugin). Skeleton AnalysisĪ second simplification is made to the image for the purpose of estimating the lengths of the midlines extracted from segmented mitochondrial structures and the extend of branching. You can check the image calibration under Image → Properties. The area or volume of the image occupied by signal is returned as the mitochondrial footprint, the units of which will depend on how the image has been calibrated. This binarized copy is overlaid upon the original image after processing as an accuracy/artifact check measure. In the binarized image, magenta represents the signal positive pixels, while the background pixels are black. Once the image has been binarized, the area or volume can be estimated by simply counting the number of signal positive pixels/voxels and multiplying by the area or volume of the pixel/voxel approximated as a rectangle or rectangular prism. If you find an issue arises using a specific thresholding method, please open an issue using the GitHub issue tracker (only a subset were tested). There are many thresholding methods available through ImageJ Ops. This is generated by automatic thresholding, a good overview of which available on the Auto Threshold page. One is a binary representation, which simply represents pixels as containing signal or being background. MiNA extracts morphological information from two simplifications of the image. Processing Pipeline and Usage Determing the Area/Volume of Mitochondria Be sure to check the closed issues as well! We may have addressed a concern already. ❓ Issue Tracking - Submit bugs or feature requests here. 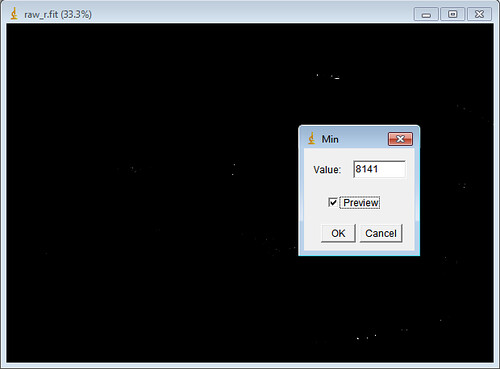
□ Source Code - Fork the project and get hacking.Look at Processing Pipeline and Usage for more up to date and thorough information. □ The Publication - The publication regarding the original macros.The links listed below provide a starting point for using, modifying, and discussing the project. It has since been altered to provide a more intuitive interface, additional options, and output. Jeff Stuart at Brock University and is based off of other analysis pipelines. The tool was developed as a result of adherent mammalian cell culture work in the laboratory of Dr. MiNA (Mitochondrial Network Analysis) is a ImageJ based utility to aid in quantitatively describing the appearance of mitochondrial morphology in fluorescence micrographs of well resolved and labelled mitochondrial structures. MiNA - ImageJ Tools for Mitochondrial Morphology Research About the Project
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
